DOP65 Mirikizumab regulates genes involved in anti-TNF resistance and ulcerative colitis disease activity
B. Steere1, J. Schmitz1, N. Powell2, R. Higgs1, K. Gottlieb1, Y. Liu1, B. Jia1, J.L. Tuttle3, W.J. Sandborn4, B.E. Sands5, V. Krishnan1
1Eli Lilly and Company, Lilly Biomedicines, Indianapolis IN, USA, 2Centre for Inflammation Biology and Cancer Immunology, King’s College London, London, UK, 3Lilly Biotechnology Center, Eli Lilly and Company, Sand Diego, CA, USA, 4Division of Gastroenterology, University California San Diego, La Jolla, CA, USA, 5Division of Gastroenterology, Icahn School of Medicine at Mount Sinai, New York, NY, USA
Background
Mirikizumab (miri), a p19-directed IL-23 antibody, demonstrated efficacy and was well-tolerated in a phase 2 randomised clinical trial in patients with moderate-to-severe UC (NCT02589665). This abstract explores gene expression changes in colonic tissue from study patients and their association with clinical outcomes.
Methods
Patients were randomised 1:1:1:1 to receive intravenous placebo (PBO,
Results
The greatest improvement in clinical outcomes at Week 12 was observed in the 200 mg miri group1; likewise, the greatest PBO-adjusted change from BL in transcripts was observed in this group. Transcripts correlating with key UC disease activity indices at BL, including modified Mayo score (MMS), ulcerative colitis Endoscopic Index of Severity (UCEIS), Geboes score, and Robarts Histopathology Index (RHI), included MMP1, MMP3, S100A8, IL1B, and UGT2A3, with the highest correlations occurring with the histopathologic indices (Figure 1). Miri treatment modulated the expression of transcriptional modules predicted to be enriched in cell profiles identified as key drivers of UC2 (Table 1, columns 1–2) as well as genes determined to be associated with UC by genome-wide association studies (GWAS; Table 1, column 3). Moreover, miri treatment affected transcripts involved in resistance to anti-TNF treatments (Table 1, column 4). A number of the genes in these categories were among those most affected by miri treatment (Table 1, columns 5–6).
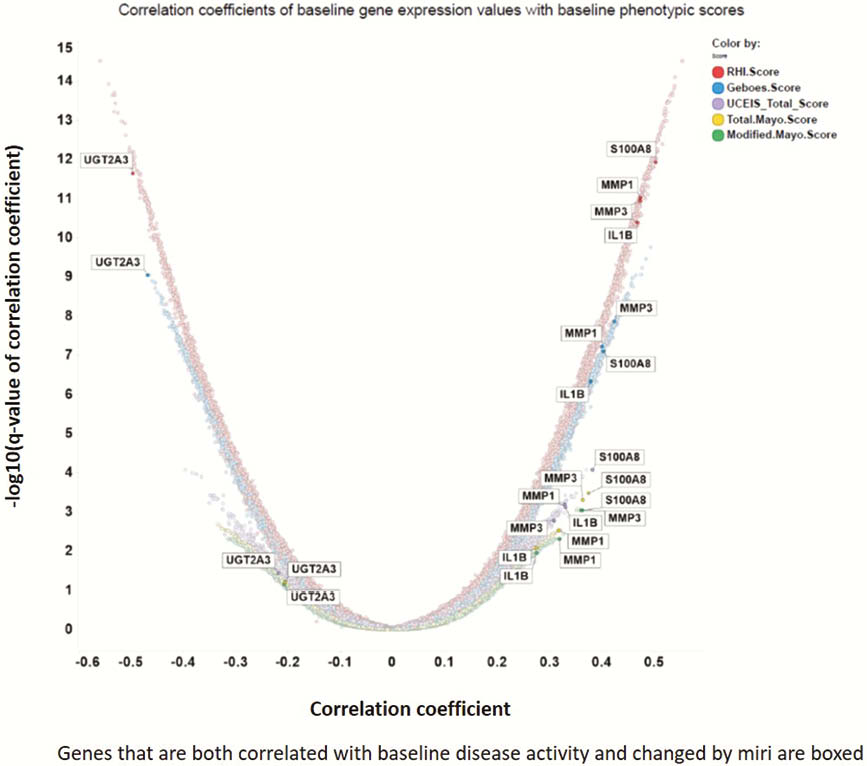
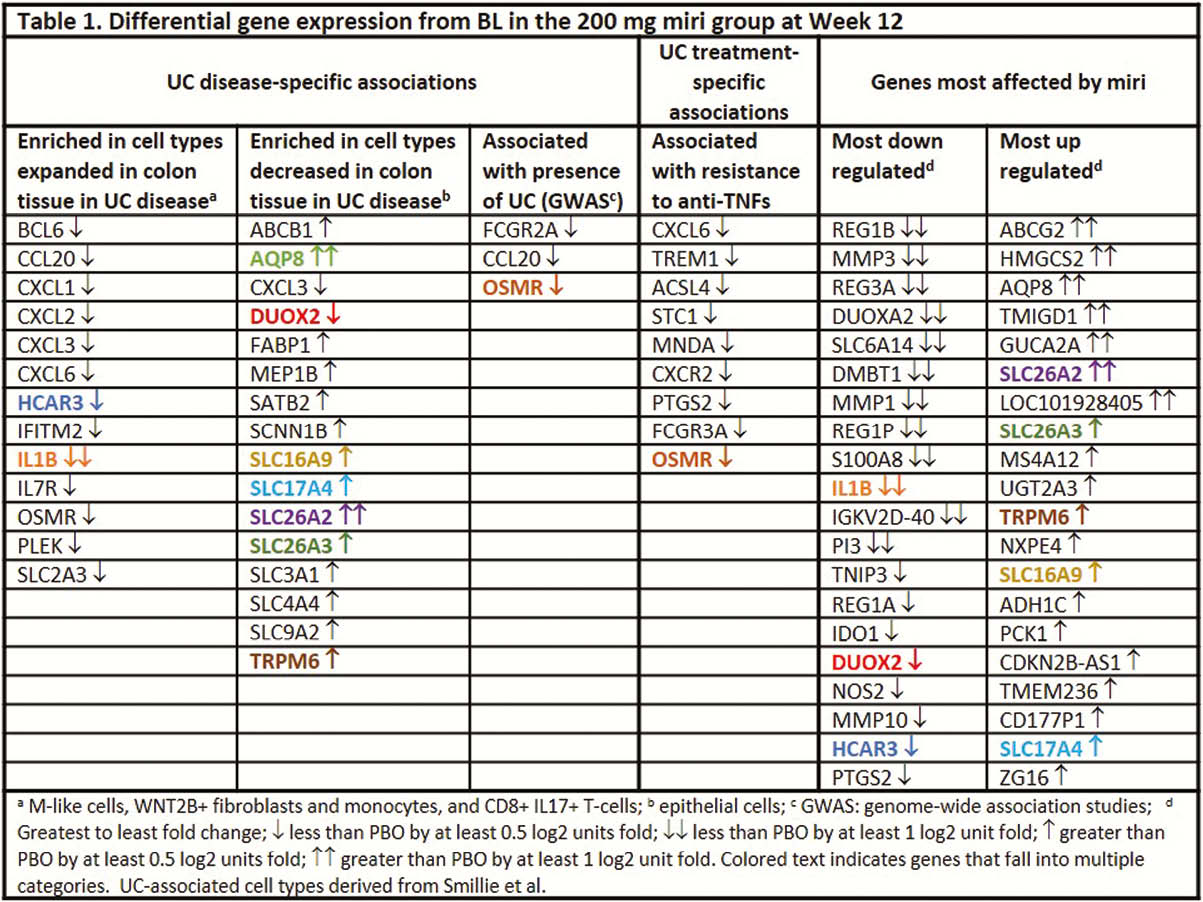
Conclusion
This is the first large-scale gene expression study of diseased tissue from UC patients treated with anti-IL23p19 therapy. It is the first study to show how anti-IL23p19 therapy modulates biological pathways involved in resistance to anti-TNFs. These results are consistent with the demonstrated efficacy of miri in patients in whom TNF antagonists have failed.
References
Sandborn WJ,
Smillie CS,


