P198 An inflammatory serum metabolomic signature predicts response to vedolizumab treatment in people with Crohn’s Disease
Radford-Smith, D.(1);Yates, A.(2);Hageman, I.(3);Joustra, V.(3);Li Yim, A.(4);Davids, M.(5);Henneman, P.(4);Hakvoort, T.(6);D'Haens, G.(3);de Jong, W.(6);Anthony, D.(1);Satsangi, J.(7);Probert, F.(2);
(1)University of Oxford, Department of Pharmacology, Oxford, United Kingdom;(2)University of Oxford, Department of Chemistry, Oxford, United Kingdom;(3)University of Amsterdam, Department of Gastroenterology and Hepatology, Amsterdam, The Netherlands;(4)University of Amsterdam, Genome Diagnostics Laboratory- Department of Clinical Genetics, Amsterdam, The Netherlands;(5)Academic Medical Center, Laboratory of Experimental Vascular Medicine, Amsterdam, The Netherlands;(6)University of Amsterdam, Tytgat Institute for Liver and Intestinal Research, Amsterdam, The Netherlands;(7)University of Oxford, Translational Gastroenterology Unit- NIHR Oxford Biomedical Research Centre, Oxford, United Kingdom;
Background
While Vedolizumab (VDZ) is an established treatment for Crohn’s disease (CD), primary non-response is well-recognised – there is an urgent clinical need for biomarkers that predict response, before initiation of treatment. Metabolomic profiling is a rapidly growing field in biomarker discovery and can distinguish between active and quiescent inflammatory bowel diseases. Here, we sought to identify a metabolic signature able to predict response to VDZ.
Methods
Prospective serum samples from 62 CD patients before (baseline) and post-treatment follow-up (FU) (median 30 weeks, IQR 27-35) were analyzed. Patients were stratified into n=35 responders (R) and n=27 non-responders (NR) using endoscopic (≥50% reduction in SES-CD score), clinical (≥3 point drop in HBI or HBI ≤4, no systemic steroids), and/or biochemical assessments (≥50% reduction in C-reactive protein (CRP) and fecal calprotectin (fCal) or a basal CRP ≤5 g/mL and fecal calprotectin ≤250 µg/g). Untargeted metabolomic profiling was carried out using high-resolution nuclear magnetic resonance (NMR) spectroscopy. Significant differences in metabolite signatures were identified by orthogonal partial least squares discriminant analysis (OPLS-DA). Model accuracy (acc.) was assessed by external 10-fold cross-validation (CV), permutation testing, and validated on an independent test set.
Results
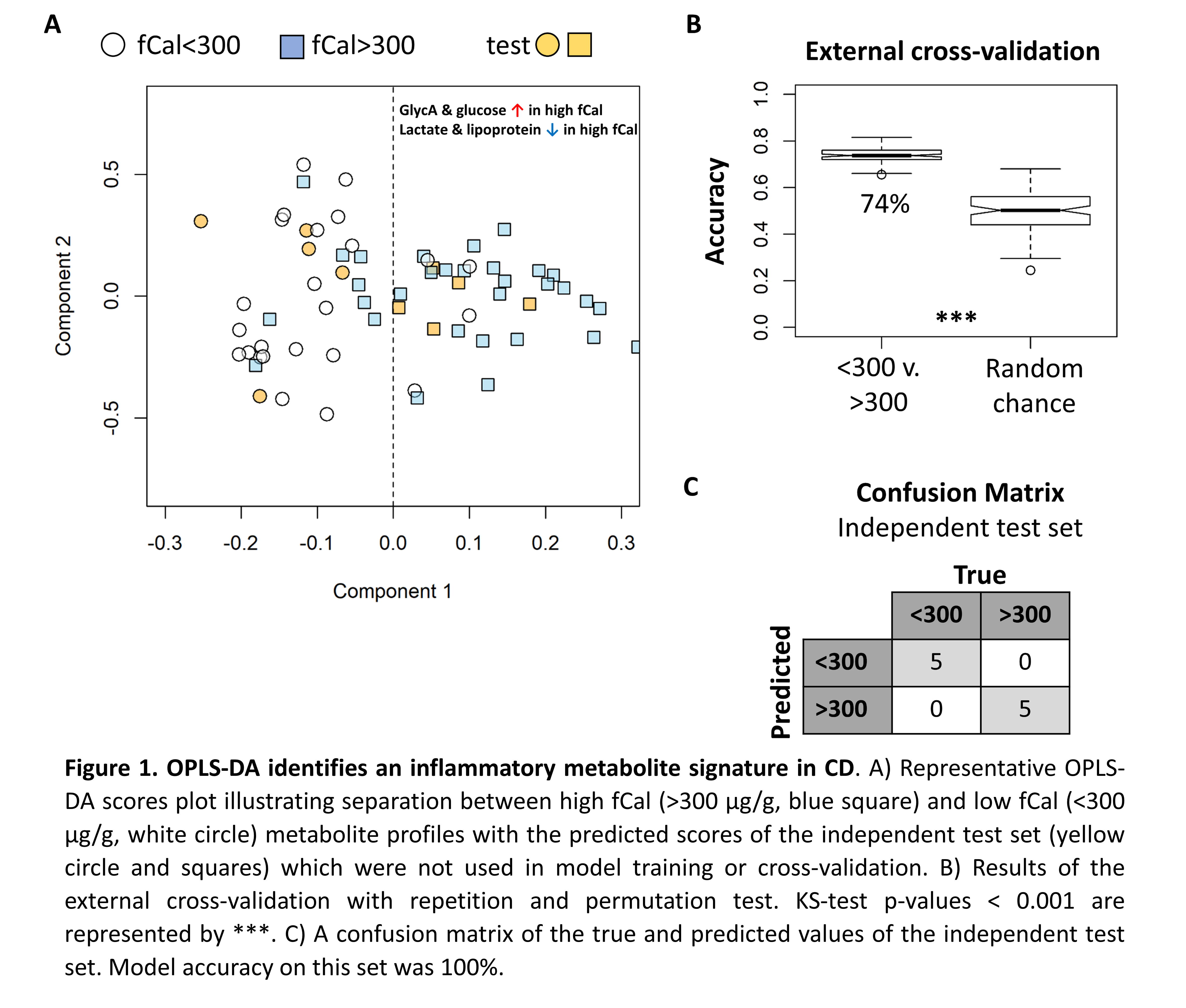
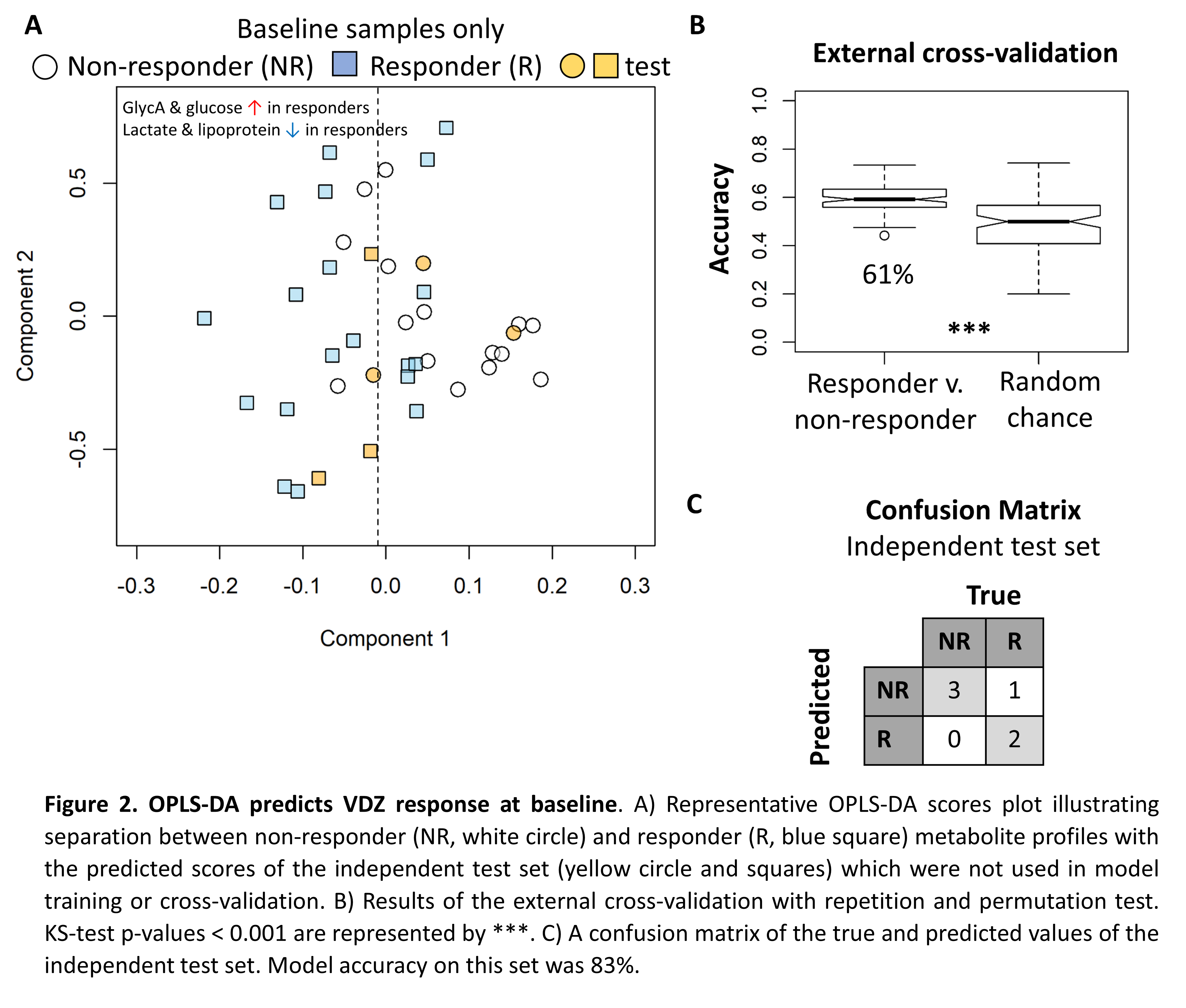
We first demonstrate that bowel preparation required for endoscopy had a significant impact on the serum metabolite profile (CV acc. 77±4%, acc. n=10 independent test set 80%, KS test p-value < 0.001) and samples taken from patients at colonoscopy were excluded from further analysis. This reduced the cohort size; 19 baseline (12R/7NR) and 25 FU (16R/9NR). No significant difference in baseline fCal or CRP was observed between R and NR (t-test p-value >0.05). The metabolite profiles at baseline and FU were indistinguishable (KS test p-value > 0.05) suggesting the metabolome was stable. However, high (>300 µg/g) fCal values were associated with elevated N-acetyl-glycoprotein (GlycA), a known inflammatory marker, elevated serum glucose along with decreased lipoprotein and lactate levels (CV acc. 74±3%, acc. n=10 independent test set 100%, KS test p-value < 0.001). This same metabolite signature predicts response (CV acc. 61±5%, acc. n=6 independent test set 83%, KS test p-value < 0.001) with responders exhibiting increased metabolic inflammation at baseline.
Conclusion
Here, we identify an inflammatory metabolic signature which predicts response to VDZ. Work to confirm these findings in a larger cohort and extend this method to investigate infliximab, adalimumab and ustekinumab treated patients, as part of the EPIC Pioneer study, is ongoing.